|
8/13/2023 0 Comments Cite sequencher
The Java heap space was exhausted forĬLUSTAL-OMEGA, FAMSA, KALIGN and MAFFT-PARTTREE were also able toĪlign the larger data set containing fragments, however, the alignment 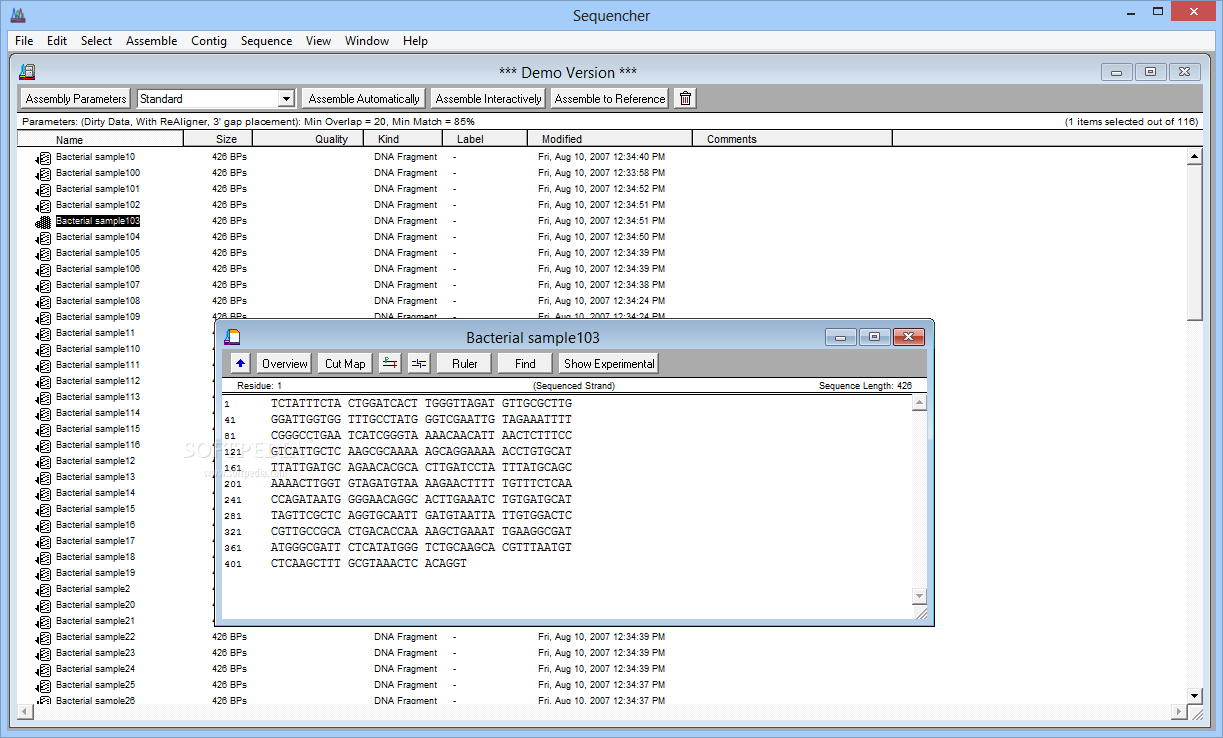
I was not able to run the full genome data set (or the largerįragmentary one), using PASTA (v1.6.3 or v1.8.2), neither on a 8Gb norĪ 256Gb nor a 512Gb machine. TheĪlignment lengths were comparable to the ones quoted by the authors.ĬLUSTAL-OMEGA(v1.2.3) T=101min RAM=1Gb L=3846 
I had to run MUSCLE on a machine with more memory. PC (8 cores, 8Gb RAM), and most of them in under 2 hours, using less Review is able to align the full genome set, most of them on a modest Each of the alignment software that I had suggested in my original However, their answers to point are highly Have responded satisfactorily to almost all the referees'Ĭriticisms. Traditional methods for phylogenetic analysis of hepatitis B viruses' The authors of 'Fully automated sequence alignment methods outperform Actually MAFFT has an option designed for adding fragmentary sequences to a backbone, as stated in the last sentence in Introduction. It seems to be a bit misleading to refer to MAFFT in the second sentence in Introduction: 'Traditionally, MSA methods for phylogenetic reconstruction are performed using alignment programs such as MUSCLE or MAFFT followed by manual, “by eye” corrections'. This might be informative since automated methods require less labour.Ģ. A possible conclusion would be that automated methods resulted in an apparently reasonable estimation for this specific data. It seems to be difficult to discuss which method is better based on these observations. * The results are not fully consistent with an external reference (genotypes classification in this case). * The results by methods A and B are not essentially different. * Estimation by method A is significantly more similar to the reference than estimation by method B. * There is a reference that can be regarded to be correct based on a solid basis and If a title says that method A outperforms method B, then readers expect: Regardless of the cause of the deviation, our results illustrate that violating key assumptions of coalescent models can mislead inferences of population history.I suggest revising the title to reflect the observations more simply. However, both selection and interspecific hybridization could account for the heterogeneity observed among loci. Defining more complex models of population history demonstrated that a pre-divergence bottleneck was also unlikely to explain this heterogeneity. Using two different coalescent methods to infer models of population history and then simulating neutral genetic diversity under these models, we found that the among-locus heterogeneity in nucleotide diversity was significantly higher than expected for these simple models. Nucleotide diversity among these loci varied by nearly two orders of magnitude (from 0.0004 to 0.029), and this heterogeneity could not be explained by differences in substitution rates. We sampled 22 nuclear intron sequences from at least 19 different chromosomes (a genomic transect) to test for deviations from selective neutrality in the gadwall (Anas strepera), a Holarctic duck. However, violating model assumptions can result in a poor fit between empirical data and the models. These inferences, such as divergence, gene flow, and changes in population size, assume that genetic data reflect simple population histories and neutral evolutionary processes. Inferring aspects of the population histories of species using coalescent analyses of non-coding nuclear DNA has grown in popularity.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
